
Additional documentation and vignettes can be found at /sctk. Specific analyses or entire workflows can be summarized and shared with comprehensive HTML reports generated by Rmarkdown.  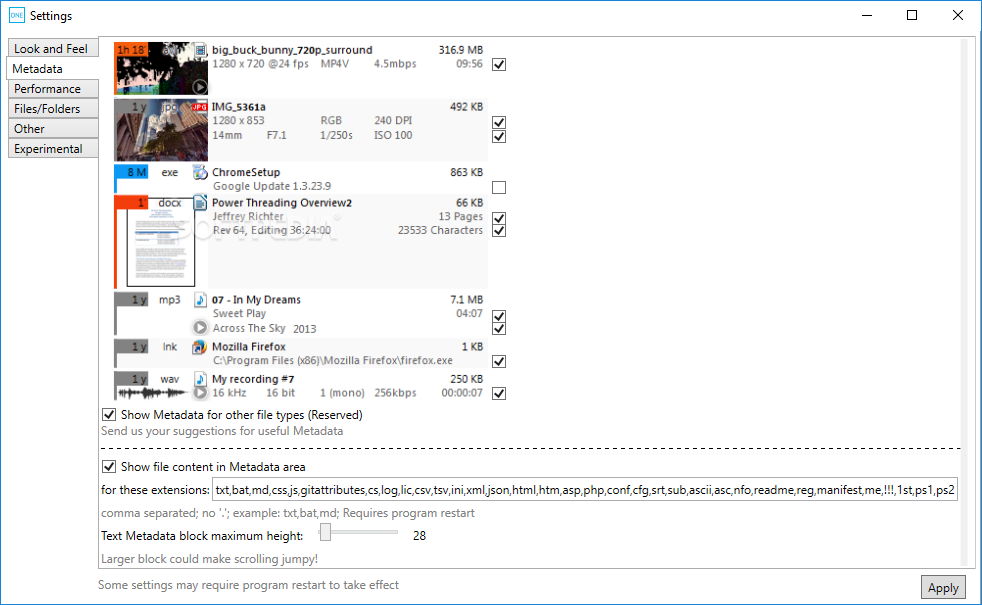
Users can analyze their data using commands in the R console or by using an interactive Shiny Graphical User Interface (GUI). Streamlined quality control can be performed on the command line using the SCTK-QC pipeline. Curated workflows can be used to run Seurat and Celda. A general "a la carte" workflow gives users the ability access to multiple methods for data importing, calculation of general QC metrics, doublet detection, ambient RNA estimation and removal, filtering, normalization, batch correction or integration, dimensionality reduction, 2-D embedding, clustering, marker detection, differential expression, cell type labeling, pathway analysis, and data exporting. SCTK allows users to seamlessly integrate tools from various packages at different stages of the analysis workflow. The Single Cell Toolkit (SCTK) in the singleCellTK package provides an interface to popular tools for importing, quality control, analysis, and visualization of single cell RNA-seq data. Comprehensive and Interactive Analysis of Single Cell RNA-Seq Data Homepage :
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
